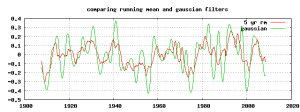
The following awk script will apply a gaussian filter to the rate of change ( derivative ) of the data. This is simpler than doing two separate operations and provides a more accurate numerical estimation of the derivative than using two point “first difference”.
Since we know the analytical form of the derivative of the gaussian function, the derivative can be applied before making a quantised approximation for numerical processing of quantised data.
Analytically, since both the convolution and the derivative are linear operations they can be done in either order and produce identical results. This allows taking the derivative of the gaussian then doing a single convolution without loss of generality.
There is still the inaccuracy of approximating the infinite gaussian function by a finite, quantised kernel but there is no loss of accuracy from also approximating the derivative. Thus the result is mathematically more accurate than using the first difference and then a gaussian filter.
The usual 3-sigma window for a gaussian filter is slightly extended to maintain similar accuracy in the D-o-G filter.
Since the derivate has high-pass properties ( attenuation inversely proportional to frequency ) and the gaussian is a low-pass filter the combined filter is a band pass filter. One common use if for edge detection, for example in medical imagery. Equally if the derivative of the data is being studied and some low-pass filtering is required it provides a one-step solution that does not rely on a digitised approximation of the derivative.
An example of edge detection is shown in determining long-term variations in the date of Arctic sea-ice minima:
Select and copy the text in the following block to get a copy of the script.
#!/bin/awk -f
# 3-sigma derviative of gaussian filter
# sigma, if not given, is 2 data points wide
# usage : ./dgauss.awk filename <sigma=2> <scale_factor=1>
# optional scale_factor simply scales the output
# sigma can be compared to the period of the -3dB point of the filter
# result is centred, ie not phase shifted. resulting dataset shorter by 3*sigma ( half window ) each end
# nov 2012 OFMT="%10.8f"
### ensure data is continuous and equally spaced ### !!
# FWHM (-3dB) = 2*sigma*sqrt(2*ln2)
# half height freq = sigma * 1.1774
# 3 sigma gives 99.7% accuracy for gauss, extend to 4-sigma for deriv-gauss
function abs(x){return (((x < 0.0) ? -x : x) + 0.0)}
BEGIN { OFMT="%10.8f"
# ARGV[1]=filename; argv[0] is script name, hence ARGC>=1
datacol=2 # fixed
if (ARGV[1]=="") { print " # usage : ./dgauss.awk filename <sigma=2> <scale_factor=1>" ; exit 1}
pi= 3.14159265359811668006
if ( ARGC >3 ) {scaleby=ARGV[3];ARGV[3]=""} else {scaleby=1};
if ( ARGC >2 ) {sigma=ARGV[2];ARGV[2]=""} else {sigma=2};
print "# filtering "ARGV[1]" with gaussian-derivative of sigma= ",sigma
root2pi_sigma=sqrt(2*pi)*sigma;
two_sig_sqr=2.0*sigma*sigma;
# extend 3 sigma window to 3.5 for derivate-gauss to maintain accuracy.
gw=(4*sigma)-1;
# calculate normalised gaussian coeffs
# for (tot_wt=j=0;j<=gw;j++) {tot_wt+=gwt[-j]=gwt[j]=exp(-j*j/two_sig_sqr)/ root2pi_sigma};
# tot_wt=0
### NB d/dt(gauss) is asymmetric, so must be time reversed to correctly do convolution.
for (tot_wt=j=0;j<=gw;j++) {tot_wt+=gwt[j]= j/sigma/sigma*exp(-j*j/two_sig_sqr) / root2pi_sigma };
tot_wt=2*tot_wt-gwt[0]; # to scale each lobe to |0.5| ; gwt[0]=0
tot_wt/=scaleby;
for (j=0;j<=gw;j++) {
# gwt[j]/=tot_wt; # gaussian is already normalised, don't rescale derivate !
gwt[-j]=-gwt[j]
};
# strip off last .xxx part of file name
# FIXME : improve this (doesn't work with paths like ../filename)
if (ARGV[1]=="-") {
out_file="-";
}
else {
split(ARGV[1],fn,".");
basename=fn[1]
out_file=basename"-gauss-deriv"sigma".dat";
print "# ",out_file >out_file;
}
ln=-1;
}
( ($0 !~ /^#/) && ($0 !~ /^[" ",\t]*$/) ) {
xdata[++ln]=$1;
ydata[ln]=$datacol;
if (ln>2*gw)
{
# cf diff.awk
# print (xdata[ln]+xdata[ln-step])/2. ,ydiff/xdiff
# print (xdata[ln]+xdata[ln-step])/2. ,ydiff/xdiff >> out_file;
xdiff=(xdata[ln]-xdata[ln-1]);
# ydiff=(ydata[ln]-ydata[ln-step]);
dgauss=0
for (j=-2*gw;j<=0;j++) {dgauss+=ydata[ln+j]*gwt[j+gw]}
if (out_file=="-")
print xdata[ln-gw],dgauss/xdiff , ydata[ln-gw]
else {
print NR,xdata[ln-gw],dgauss/xdiff ,ydata[ln-gw]
print xdata[ln-gw],dgauss/xdiff , ydata[ln-gw] >> out_file;
}
}
else
{
# print $1,$2;
}
}
END {
if (ARGV[1]=="") { exit 1}
print "# gausssian-derivative window width = "gw+gw+1",done"
print "# output file = "out_file
# sum=0; for (j=-gw;j<=gw;j++) {sum+= abs(gwt[j]);}
# print "# abs sum of coeffs = ",sum;
}